Research
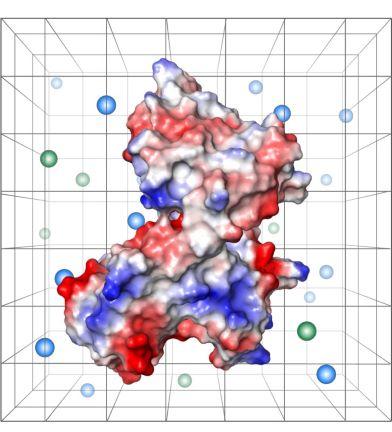
Our research group, based in the Department of Surgery in Cambridge, is focused on interdisciplinary and translational research that aims to increase access to transplantation and to improve long-term transplant outcomes. We are particularly interested in the human major histocompatibility complex proteins (called Human Leukocyte Antigens or HLA) and their role in transplant allo-immunity and graft injury. Much of our efforts in this area are focused on studying the genetic and structural characteristics of HLA proteins and the functional properties of HLA-specific antibodies to assess transplant immunological risk and to improve donor-recipient tissue matching. We are also interested in big-data analysis for phenotyping and forecasting in transplantation and in applying technological innovations, such as machine perfusion, to assess the quality of donor organs and improve their function after transplantation.
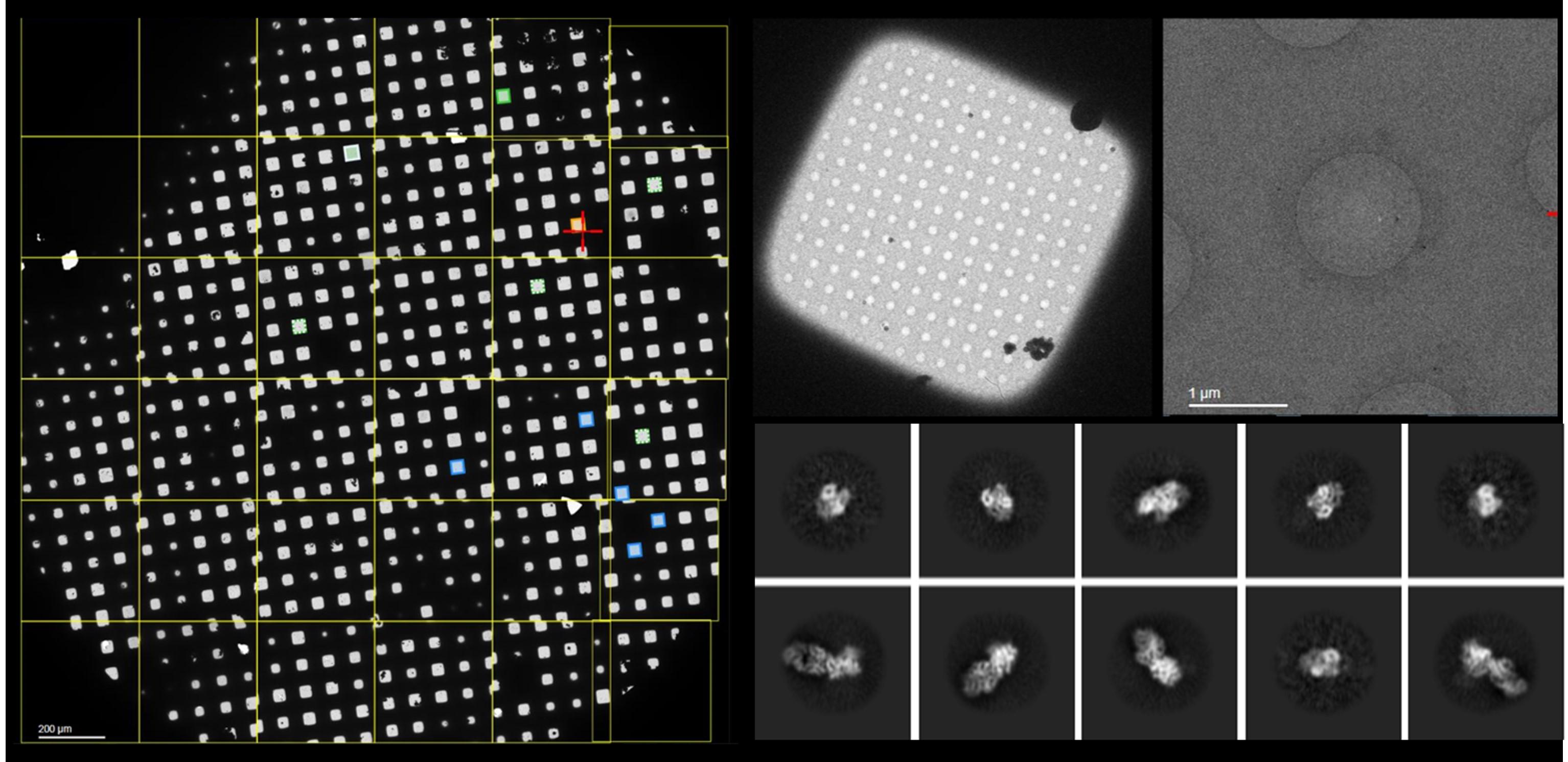
We work collaboratively with colleagues in academia and industry and utilise a variety of techniques to address our research questions, including computational structural and physicochemical protein modelling, cryo-Electron microscopy, protein chemistry, molecular biology methods, microfluidics, transcriptomics and proteomics analyses. Research in the group has been licensed to industry and led to the development of widely used HLA immunogenicity analysis software.
Selected Publications
Schneider MM*, Scheidt T*, Priddey AJ*, Xu CK*, Hu M, Meisl G, Devenish SRA, Dobson CM, Kosmoliaptsis V*, Knowles TPJ*. Microfluidic antibody affinity profiling of alloantibody-HLA interactions in human serum. Biosensors and Bioelectronics 2023; 228: 115196
Isaacson D, Schold JD, Gmeiner MW,Copley HC,Kosmoliaptsis V*, Tambur AR*. HLA-DQ Mismatches Lead to More Unacceptable Antigens, Greater Sensitization, and Increased Disparities in Repeat Transplant Candidates. JASN 2022; 33: 2293
Wu X, Senanayake R, Goodchild E, et al. [11C]metomidate PET-CT versus adrenal vein sampling for diagnosing surgically curable primary aldosteronism: a prospective, within-patient trial. Nature Medicine 2023; 29: 190-202
Denninger V, Xu CK, Meisl G, Morgunov AS, Fiedler S, Ilsley A, Emmenegger M, Malik AY, Piziorska MA, Schneider MM, Devenish SRA, Kosmoliaptsis V, Aguzzi A, Fiegler H, Knowles TPJ. Microfluidic Antibody Affinity Profiling Reveals the Role of Memory Reactivation and Cross-Reactivity in the Defense Against SARS-CoV-2. ACS Infect Dis. 2022; 8: 790
Gaurav R, Butler AJ, Kosmoliaptsis V, Mumford L, Fear C, Swift L, Fedotovs A, Upponi S, Khwaja S, Richards J, Allison M, Watson CJE. Liver Transplantation Outcomes From Controlled Circulatory Death Donors: SCS vs in situ NRP vs ex situ NMP. Annals of Surgery 2022; 275: 1156
Tambur AR, Kosmoliaptsis V, Claas FHJ, Mannon RB, Nickerson P, Naesens M. Significance of HLA-DQ in kidney transplantation: time to reevaluate human leukocyte antigen-matching priorities to improve transplant outcomes? An expert review and recommendations. Kidney Int. 2021; 100: 1012
Wiebe C, Kosmoliaptsis V, Pochinco D, Gibson IW, Ho J, Birk PE, Goldberg A, Karpinski M, Shaw J, Rush DN, Nickerson PW. HLA-DR/DQ molecular mismatch: A prognostic biomarker for primary alloimmunity. Am J Transplant. 2019; 19: 1708
Mallon DH, Kling C, Robb M, Ellinghaus E, Bradley JA, Taylor CJ, Kabelitz D, Kosmoliaptsis V. Predicting Humoral Alloimmunity from Differences in Donor and Recipient HLA Surface Electrostatic Potential. J Immunol. 2018; 201: 3780
Watson CJE, Kosmoliaptsis V, Pley C, Randle L, Fear C, Crick K, Gimson AE, Allison M, Upponi S, Brais R, Jochmans I, Butler AJ. Observations on the ex situ perfusion of livers for transplantation. Am J Transplant. 2018; 18: 2005
Kosmoliaptsis V, Mallon DH, Chen Y, Bolton EM, Bradley JA, Taylor CJ. Alloantibody Responses After Renal Transplant Failure Can Be Better Predicted by Donor-Recipient HLA Amino Acid Sequence and Physicochemical Disparities Than Conventional HLA Matching. Am J Transplant. 2016; 16: 2139
Goyette P, Boucher G, Mallon D, Ellinghaus E, Jostins L, Huang H, Ripke S, Gusareva ES, Annese V, Hauser SL, Oksenberg JR, Thomsen I, Leslie S; International Inflammatory Bowel Disease Genetics Consortium; Australia and New Zealand IBDGC; Belgium IBD Genetics Consortium; Italian Group for IBD Genetic Consortium; NIDDK Inflammatory Bowel Disease Genetics Consortium; United Kingdom IBDGC; Wellcome Trust Case Control Consortium; Quebec IBD Genetics Consortium, Daly MJ, Van Steen K, Duerr RH, Barrett JC, McGovern DP, Schumm LP, Traherne JA, Carrington MN, Kosmoliaptsis V, Karlsen TH, Franke A, Rioux JD. High-density mapping of the MHC identifies a shared role for HLA-DRB1*01:03 in inflammatory bowel diseases and heterozygous advantage in ulcerative colitis. Nature Genetics 2015; 47: 172